Research Areas
Bioimage Informatics
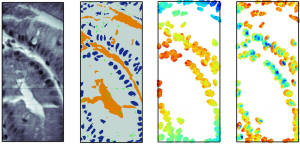 Bioimage Informatics draws upon advances in signal processing, optics, probe chemistry, molecular biology and machine learning to provide answers to biological questions from the growing numbers of biological images acquired in digital form. Microscopy is one of the oldest biological methods, and for centuries it has been paired with visual interpretation to learn about biological phenomena. With the advent of sensitive digital cameras and the dramatic increase in computer processing speeds over the past two decades, it has become increasingly common to collect large volumes of biological image data that create a need for sophisticated image processing and analysis. In addition, dramatic advances in machine learning during the same period set the stage for converting imaging from an observational to a computational discipline and allow the direct generation of biological knowledge from images.
Bioimage Informatics draws upon advances in signal processing, optics, probe chemistry, molecular biology and machine learning to provide answers to biological questions from the growing numbers of biological images acquired in digital form. Microscopy is one of the oldest biological methods, and for centuries it has been paired with visual interpretation to learn about biological phenomena. With the advent of sensitive digital cameras and the dramatic increase in computer processing speeds over the past two decades, it has become increasingly common to collect large volumes of biological image data that create a need for sophisticated image processing and analysis. In addition, dramatic advances in machine learning during the same period set the stage for converting imaging from an observational to a computational discipline and allow the direct generation of biological knowledge from images.
Cellular and Systems Modeling
Cellular and Systems Modeling undertakes the ambitious task of studying the dynamics of biological 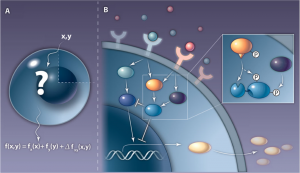 and biomedical processes from a whole system point of view. The observed systems range over orders of magnitude, from tissue to cells to molecular assemblies! Engineering tools are used along with genome-scale information in mathematical and/or computational models that usually adopt a top-down approach. Modeling diseases, entire ‘virtual’ cells, or subcellular networks of interactions are among typical tasks. Major research topics include the modeling of complex signaling and regulatory networks, transport mechanisms, spatio-temporal evolution of microphysiological events, as well as establishing the links between the development of complex phenotypes and the seemingly unrelated molecular events.
and biomedical processes from a whole system point of view. The observed systems range over orders of magnitude, from tissue to cells to molecular assemblies! Engineering tools are used along with genome-scale information in mathematical and/or computational models that usually adopt a top-down approach. Modeling diseases, entire ‘virtual’ cells, or subcellular networks of interactions are among typical tasks. Major research topics include the modeling of complex signaling and regulatory networks, transport mechanisms, spatio-temporal evolution of microphysiological events, as well as establishing the links between the development of complex phenotypes and the seemingly unrelated molecular events.
Computational Genomics
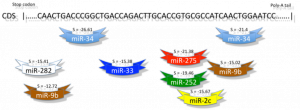 Computational Genomics entails efforts to digest the daunting quantity of genomic and proteomic data now available by systematic development and application of probability and statistics theories, information technologies and data mining techniques. Linguistics methods are viewed as promising tools towards elucidating sequence-structure-function relations, and complementing computational genomics studies. Computational genomics targets understanding gene/protein function, identifying and characterizing cellular regulatory networks and discerning the link between genes and diseases. Discovery and processing of this information is pivotal in the development of novel gene therapy strategies and tools.
Computational Genomics entails efforts to digest the daunting quantity of genomic and proteomic data now available by systematic development and application of probability and statistics theories, information technologies and data mining techniques. Linguistics methods are viewed as promising tools towards elucidating sequence-structure-function relations, and complementing computational genomics studies. Computational genomics targets understanding gene/protein function, identifying and characterizing cellular regulatory networks and discerning the link between genes and diseases. Discovery and processing of this information is pivotal in the development of novel gene therapy strategies and tools.
Biological Physics
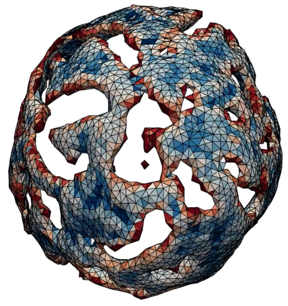 Biological physics encompasses a multidisciplinary approach that uses principles from physics to gain insights into the fundamental processes underlying living systems. Concepts from statistical physics, dynamical systems, and fluid dynamics are applied to investigate phenomena such as cell state transitions, cell motility, tissue morphogenesis, and evolution. Biological physicists probe these multi-scale phenomena using approaches from theoretical analyses, quantitative modeling, machine learning, and experimental measurements of forces and fields. This field aims to unravel the intricate workings of life from a unique perspective, which can also lead to new discoveries in physics and biology.
Biological physics encompasses a multidisciplinary approach that uses principles from physics to gain insights into the fundamental processes underlying living systems. Concepts from statistical physics, dynamical systems, and fluid dynamics are applied to investigate phenomena such as cell state transitions, cell motility, tissue morphogenesis, and evolution. Biological physicists probe these multi-scale phenomena using approaches from theoretical analyses, quantitative modeling, machine learning, and experimental measurements of forces and fields. This field aims to unravel the intricate workings of life from a unique perspective, which can also lead to new discoveries in physics and biology.
Computational Structural Biology
Computational Structural Biology aims at establishing biomolecular sequence-structure-function relations 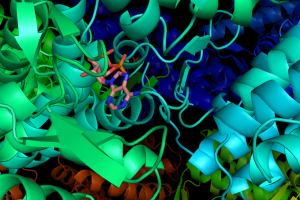 using fundamental principles of physical sciences in theoretical models and simulations of structure and dynamics. After the advances in complete genomes sequencing, it became evident that structural information is needed for understanding the origin and mechanisms of biological interactions, and designing/controlling function. Computational Structural Biology emerged as a tool for efficient identification of structure and dynamics in many applications. Major research topics include protein folding, protein dynamics with emphasis on large complexes and assemblies, protein-protein, protein-ligand and protein-DNA interactions and their functional implications. Drug design and protein engineering represent applications of note.
using fundamental principles of physical sciences in theoretical models and simulations of structure and dynamics. After the advances in complete genomes sequencing, it became evident that structural information is needed for understanding the origin and mechanisms of biological interactions, and designing/controlling function. Computational Structural Biology emerged as a tool for efficient identification of structure and dynamics in many applications. Major research topics include protein folding, protein dynamics with emphasis on large complexes and assemblies, protein-protein, protein-ligand and protein-DNA interactions and their functional implications. Drug design and protein engineering represent applications of note.